Electronic Health Records (EHRs) have revolutionized healthcare and biomedical research over the past three decades. In recent years, with the advent of new hardware and storage capabilities, EHRs have evolved into a complex framework, housing massive amounts of digital data. In providing a snapshot of a patient’s state of health, EHRs have created new opportunities to investigate the predictors and properties of health-related events across large and heterogeneous populations, which can provide clarity and insights into the future effects of medical decisions. At the individual level, these trajectories are becoming the basis for personalized medicine. Across patient cohorts, EHRs and derived trajectories provide a vital resource to understand population health management and make better decisions for healthcare operational policies1.
Given a specific disease, heterogeneity among patients usually leads to different progression patterns and may require different types of interventions, despite equivalence at the diagnostic level. This is particularly evident for complex disorders, whose disease etiology is still mostly unknown, possibly due to multiple genetic, environmental, and lifestyle factors. Patients with complex disorders may differ on multiple levels of analyses (e.g., different clinical measures such as laboratory tests or medications, or different comorbidities) and in response to treatments throughout the disease trajectory, making these conditions difficult to evaluate. Several different conditions have been referred to as complex, such as Parkinson’s disease (PD)2, multiple myeloma (MM)3, and type 2 diabetes (T2D)4.
EHR histories offer a way to examine disease complexity and present an opportunity to refine diseases into subtypes and tailor personalized treatments. This task is usually referred to as "EHR-based patient stratification" and from a computational perspective, it is a data-driven, unsupervised learning task that groups patients according to their clinical characteristics5. Deep learning has been recently applied to derive more robust patient representations to improve disease subtyping5, 6. Such approach, however, usually focuses on curated and small disease-specific cohorts, with ad hoc manually selected features. This not only limits scalability and generalizability, but also hinders the possibility to discover unknown patterns that might characterize a condition.
Our work proposes a general framework for identifying disease subtypes at scale (see Fig. 1a). We first combined an unsupervised deep learning architecture (ConvAE) to derive vector-based patient representations from a large and domain-free collection of EHRs (see Fig. 1b), then we showed that the representations learned from real-world EHRs of ~1.6M patients from the Mount Sinai Health System in New York improve clustering of patients with different disorders compared to several commonly used baselines. Last, we used the encodings learned from domain-free and heterogeneous EHRs to derive subtypes for different complex disorders and provide a qualitative analysis to determine their clinical relevance.

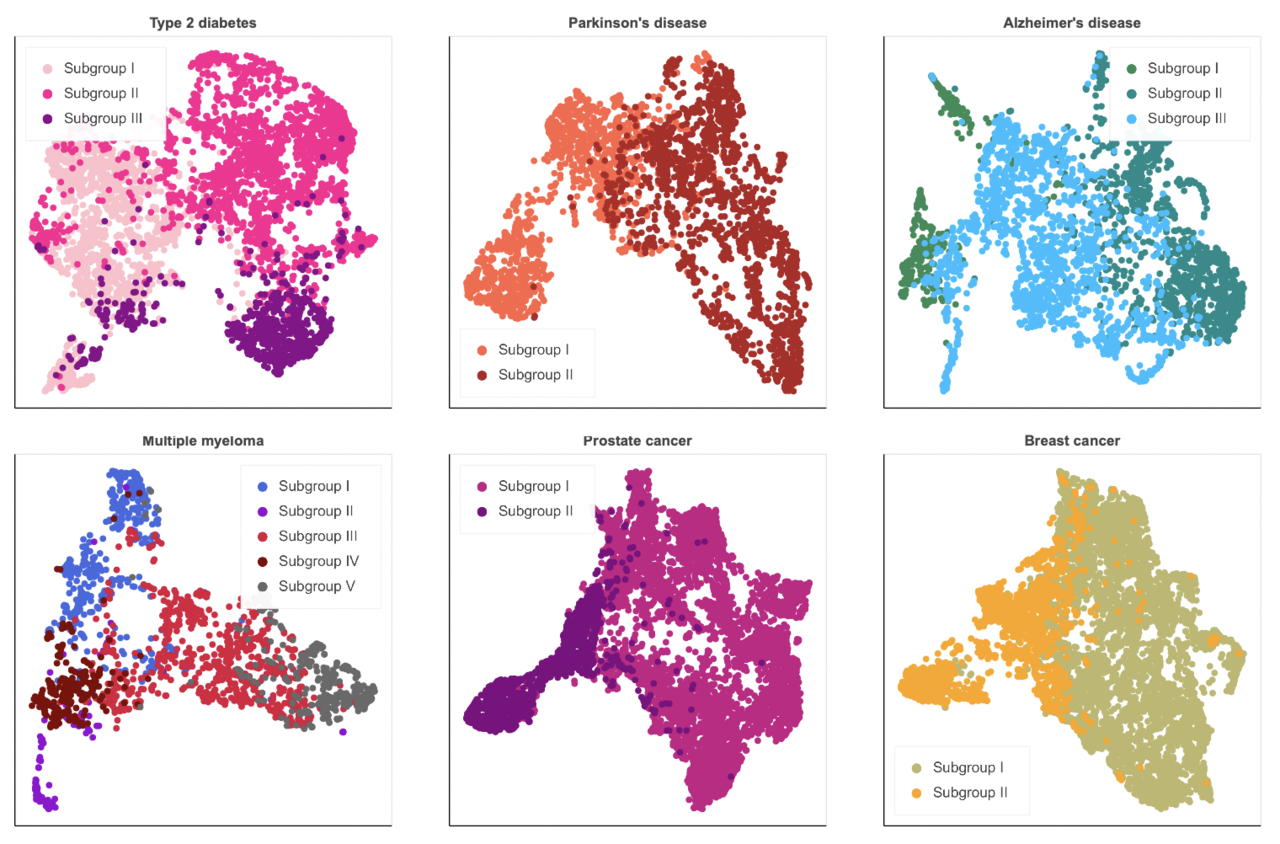
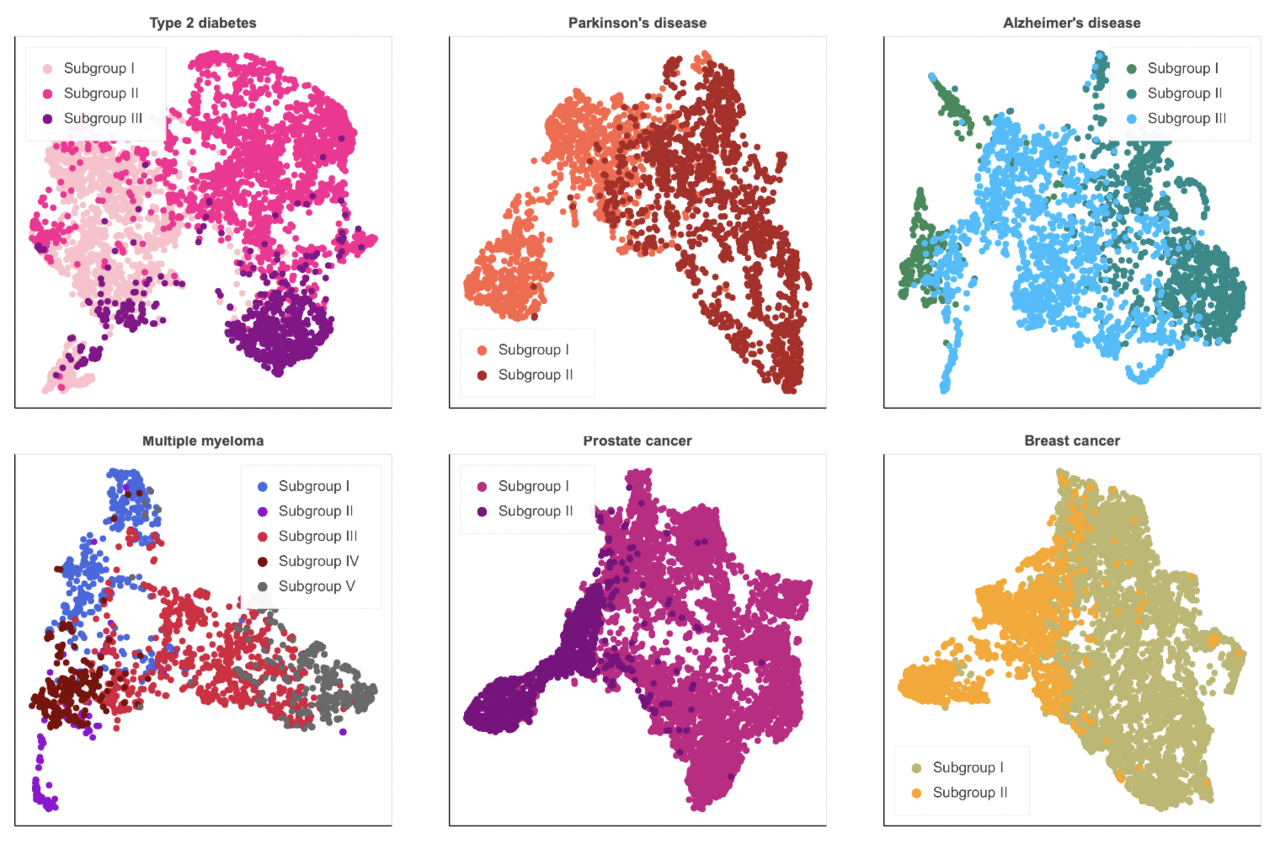
ConvAE-learned representations outperformed common baseline encodings in detecting eight complex disorders (i.e., PD, Alzheimer’s disease - AD, MM, T2D, prostate cancer - PC, breast cancer - BC, attention deficit hyperactivity disorder, and Crohn’s disease). Moreover, we were able to determine that the same paradigm allows us to detect clinically meaningful subtypes within six of these high-level disease designations, namely T2D, PD, AD, MM, PC, BC. What emerged is that sex may play a central role in the heterogeneous manifestations of these conditions (e.g., Alzheimer’s disease) and should be taken into account in stratification studies. Second, disease progression, symptom severity, and comorbidities seem to contribute the most to the phenotypic variability of complex disorders. Patients with T2D divides into three subgroups according to comorbidities (i.e., cardiovascular and microvascular problems) and symptom severity (i.e. newly diagnosed with milder symptoms). Individuals with PD show different disease durations and symptoms (i.e., motor, nonmotor). AD profiles distinguish early- and late-onset groups and separate patients with mild neuropsychiatric symptoms and cerebrovascular disease from patients with mild-to-moderate dementia. Patients with MM are characterized by different comorbidities (e.g., amyloidosis, pulmonary diseases) that manifest alongside precise typical signs of MM. Patients with PC and BC separate according to disease progression.
By learning from a large EHR that includes a diverse and heterogeneous population, our method presents opportunity for both generalizability and scalability in the identification and categorization of disease patterns using EHR specific data. Applied to complex disorders, this architecture can be leveraged to detect disease subtypes that reflect disease heterogeneity within patient populations. Moreover, if used to infer latent representations of new patients, ConvAE model can contribute to derive the subtype of any particular patient for any particular disease, and track subsequent disease progression and treatment effectiveness.
References
- Jensen, P. B., Jensen, L. J. & Brunak, S. Mining electronic health records: towards better research applications and clinical care. Rev. Genet.13, 395 (2012).
- Langston, J. W. The Parkinson’s complex: Parkinsonism is just the tip of the iceberg. Neurol.59, 591–596 (2006).
- de Mel, S., Lim, S. H., Tung, M. L. & Chng, W. J. Implications of heterogeneity in multiple myeloma. BioMed Res. Int.1–12, https://doi.org/10.1155/2014/232546 (2014).
- Pearson, E. R. Type 2 diabetes: a multifaceted disease. Diabetologia62, 1107–1112 (2019).
- Baytas, I. M. et al. Patient subtyping via time-aware LSTM Networks. In 23rd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining(eds Matwin S, S., Yu, S. & Farooq, F.) 65–74 (ACM, New York, 2017).
- Zhang, X. et al. Data-driven subtyping of Parkinson’s disease using longitudinal clinical records: a cohort study. Scientific Rep.9, 797 (2019).






Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in